How the “Swine” flu virus develops drug resistance
A University of Bristol team used the Emerald GPU supercomputer to investigate how mutations of a key enzyme of H1N1 “Swine” flu lead to the development of resistance to current antiviral flu treatments.
H1N1-2009 is a highly adaptive virus derived from different gene segments of swine, avian and human influenza. Within months of its appearance in early 2009, the H1N1-2009 strain caused the first flu pandemic of the 21st-century. The antiviral drugs zanamivir (Relenza) and oseltamivir (Tamiflu), which target the neuraminidase enzyme in influenza, successfully treated the infection but widespread use of these drugs led to a series of mutations in neuraminidase that reduce the drugs’ effectiveness. Clinical studies indicate that the mutant of swine flu neuraminidase known as IRHY reduced the effectiveness of zanamivir by 21 times and oseltamivir by 12,374 times – that is, to the point where it has become an ineffective treatment.
To understand why the effectiveness of these drugs is so seriously reduced by the occurrence of this mutation, a team of scientists at the University of Bristol, led by Prof. Adrian Mulholland and Dr. Christopher Woods, used the Emerald GPU supercomputer to perform long-timescale molecular dynamics simulations of different mutants of neuraminidase. The Bristol scientists were able to exhaustively investigate how different mutations of neuraminidase affect the binding of all of the commercially and experimentally available antiviral treatments of flu. 108 of Emerald’s 374 GPUs were used to generate 18,000 nanoseconds of molecular dynamics trajectories of influenza. Emerald generated 400 nanoseconds per day, completing the entire simulation in just six weeks. This is incredibly fast, beyond anything else available in the UK at the time. This same simulation would take three months to complete if the entire University of Bristol supercomputer, Bluecrystal was used. This is amazing, especially when you consider that Bluecrystal was the 86th fastest supercomputer in the world in 2008.
Early results provided the scientists with a deep new insight into the mechanisms by which mutations give influenza resistance to antiviral treatments. It is known experimentally that neuraminidase can exist in two conformations; an “open” form and a “closed” form. Current antiviral drugs bind only to the closed form of neuraminidase. Simulations have revealed that the drugs do indeed bind to the closed form, and keep neuraminidase closed by making strong hydrogen bonds that hold the structurally important 150-loop into the closed conformation. Simulations on Emerald revealed how mutations of the enzyme lead to disruption of the hydrogen bonds that pin close the 150-loop. This leads to this loop opening, allowing neuraminidase to change from the closed to the open conformation. The antiviral drugs bind weakly with the open conformation, thus rendering them ineffective. These findings suggest that drug resistance could be overcome by designing new antiviral drugs that make stronger hydrogen bonds between the drug and residues in the 150-loop, with the aim of helping them hold onto the 150-loop, thereby locking neuraminidase into the closed conformation.
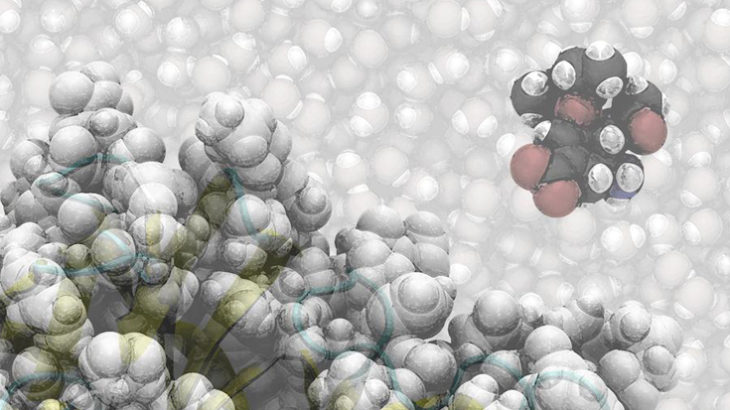
Oseltamivir (Tamiflu) approaches the binding site of neuraminidase, a key enzyme of the influenza virus. Mutations of this enzyme are stopping existing influenza drugs from binding, rendering them ineffective treatments for flu.
Contact

Professor Adrian Mulholland
School of Chemistry

